Affinity#
- class torchdr.Affinity(metric: str = 'sqeuclidean', zero_diag: bool = True, device: str = 'auto', backend: str | FaissConfig | None = None, verbose: bool = False, random_state: float | None = None, compile: bool = False, _pre_processed: bool = False)[source]#
-
Base class for affinity matrices.
- Parameters:
metric (str, optional) – Distance metric for pairwise distances. Default is “sqeuclidean”.
zero_diag (bool, optional) – Whether to set the diagonal to zero. Default is True.
device (str, optional) – Device for computation.
"auto"uses the input data’s device. Default is “auto”.backend ({"keops", "faiss", None} or FaissConfig, optional) – Backend for handling sparsity and memory efficiency. Default is None (standard PyTorch).
verbose (bool, optional) – Verbosity. Default is False.
compile (bool, optional) – Whether to compile the affinity computation. Default is False.
_pre_processed (bool, optional) – If True, skips
to_torchconversion (inputs are already tensors on the correct device). Default is False.
Examples using Affinity:#
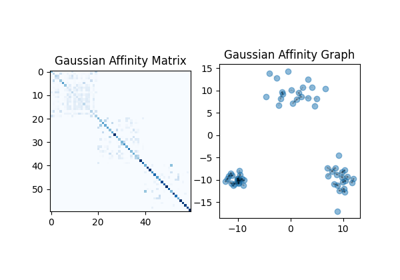
Entropic Affinities can adapt to varying noise levels

Neighbor Embedding on genomics & equivalent affinity matcher formulation